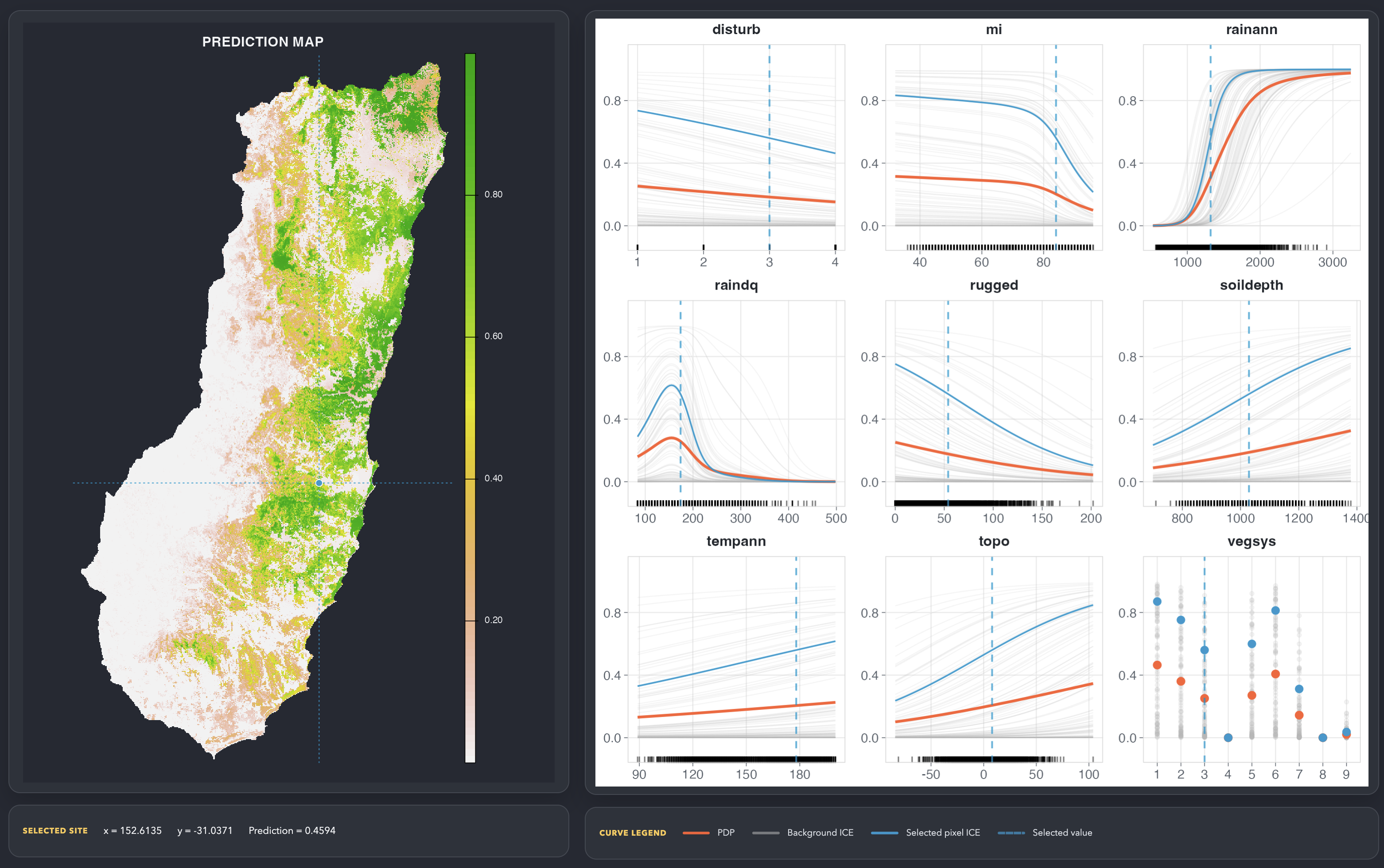
curves is an experimental R package for plotting response curves from fitted models with ggplot2. The package is intentionally small and model-agnostic: supply a fitted model, predictor data, and, when needed, a custom prediction function.

The figure above shows partial dependence curves from the included species distribution vignette.
The current API is centred around five exported functions:
univariate() for one-predictor response curves with
method = "profile", "pdp", "ice",
"ice+pdp", or "ale".bivariate() for two-predictor profile, PDP, or ALE
surfaces as static heatmaps, filled contours, or interactive 3D
plotly surfaces.interactions() for ranking numeric predictor pairs by
the strength of their centred second-order ALE interaction
surfaces.multimodel() for ensemble profile, PDP, or ALE curves
across multiple fitted models, with optional interval ribbons and
member-model overlays.mapcurve() for Shiny-based exploration that links a
prediction raster to response curves at clicked map cells.A few practical details are worth calling out:
terra::SpatRaster objects.predict() returns multiple columns,
response can be used to choose the column to plot.ggplot2 objects, so they can be
styled or combined in downstream workflows.In short, profile curves use one reference row, PDP averages predictions over sampled rows, ICE keeps those row-level curves visible, and ALE accumulates local prediction differences within the observed predictor distribution.
curves is not on CRAN. Install the development version
from GitHub:
install.packages("remotes")
remotes::install_github("rvalavi/curves")Optional packages:
terra for raster-backed predictor inputs.plotly for interactive 3D surfaces.mgcv for the GAM ensemble example below.randomForest and disdat for the species
distribution vignette.library(curves)
model <- lm(
Sepal.Length ~ Sepal.Width + Petal.Length + Petal.Width,
data = iris
)
predictors <- iris[, c("Sepal.Width", "Petal.Length", "Petal.Width")]
# Partial dependence curves (default)
univariate(model, predictors)
# Single-profile response curves
univariate(
model,
predictors,
method = "profile"
)
# Accumulated local effects curves
univariate(model, predictors, method = "ale", n = 40)
# Bivariate response surface
bivariate(
model,
predictors,
pairs = c("Sepal.Width", "Petal.Length"),
background_n = 50,
rug = TRUE,
plot_type = "heatmap"
)
# Rank pairwise ALE interactions
interactions(model, predictors, n = 10)
# Plot only the strongest ALE surfaces
bivariate(model, predictors, method = "ale", top_n = 3, n = 10)For ensemble modelling or repeated-fit comparisons, pass a list of
fitted models to multimodel(). This is useful not only for
formal ensembles, but also for cross-validation folds, bootstrap or
bagged refits, and models fit with different background samples or
closely related training sets. The models should share compatible
predictors, and their predictions should be on the same response scale.
If a set of models shares the same prediction interface, pass
non-default prediction arguments through .... For mixed
model types, supply fun as a list of wrappers, one per
model. If a shared prediction function returns multiple prediction
columns, either set response or provide a small wrapper
through fun. Use agg, weights,
interval, and show_models to control how the
model curves are combined and displayed.
models <- list(
lm(Sepal.Length ~ Sepal.Width + Petal.Length, data = iris),
mgcv::gam(Sepal.Length ~ s(Sepal.Width) + s(Petal.Length), data = iris)
)
multimodel(
models,
predictors[, c("Sepal.Width", "Petal.Length")],
background_n = 200,
show_models = TRUE
)The package includes a fuller species distribution vignette built around a down-sampled random forest classifier. It demonstrates:
response = "1"univariate()You can open it after installation with:
vignette("random-forest-species-distribution", package = "curves")mapcurve() opens a Shiny explorer that links a predicted
map to fitted response curves. Clicking a raster cell marks that site’s
predictor values on the curve panels, which helps compare local
conditions with profile, PDP, ICE, or ALE summaries.