forrest creates publication-ready forest plots from any
data frame that contains point estimates and confidence intervals —
regression models, subgroup analyses, meta-analyses, dose-response
patterns, and more. A single dependency (tinyplot) keeps
the footprint minimal.
Key features:
section argument) — no manual NA rowssubsection for nested grouping
structuresis_summary)" (Ref.)"stripe = TRUE)section_colssave_forrest()data.frame, tibble, and
data.tableinstall.packages("forrest")
# Development version from GitHub:
# install.packages("pak")
pak::pak("lorenzoFabbri/forrest")The minimal call requires only three column names —
estimate, lower, and upper. Use
group and dodge = TRUE to place multiple CIs
per row when comparing results across categories (e.g., time periods,
sexes, models). See the Introduction
vignette for the full feature tour.
library(forrest)
# Two estimates per exposure (e.g., two time windows)
periods <- data.frame(
exposure = rep(c("Metals", "OPPs", "PFAS", "Phenols", "Phthalates"), each = 2),
period = rep(c("Pregnancy", "Childhood"), 5),
est = c( 0.03, -0.04, -0.03, 0.04, -0.17, -0.02, 0.10, -0.02, 0.07, 0.10),
lo = c(-0.25, -0.26, -0.19, -0.12, -0.40, -0.24, -0.06, -0.10, -0.15, -0.08),
hi = c( 0.30, 0.19, 0.13, 0.21, 0.06, 0.19, 0.26, 0.07, 0.29, 0.28)
)
forrest(
periods,
estimate = "est",
lower = "lo",
upper = "hi",
label = "exposure",
group = "period",
dodge = TRUE,
header = "Exposure class",
ref_line = 0,
xlab = "Coefficient (95% CI)"
)
Use log_scale = TRUE and ref_line = 1 for
ratio measures, is_summary = TRUE to draw the pooled row as
a filled diamond, and weight to scale point sizes by
inverse-variance weights. See the Meta-analysis
vignette for subgroup analyses and multi-exposure layouts.
ma <- data.frame(
study = c("Adams 2016", "Baker 2018", "Chen 2019",
"Davis 2020", "Evans 2021", "Pooled"),
est = c(1.45, 0.88, 1.21, 1.07, 0.95, 1.09),
lo = c(1.10, 0.65, 0.92, 0.80, 0.72, 0.94),
hi = c(1.91, 1.19, 1.59, 1.43, 1.25, 1.26),
weight = c(180, 95, 140, 210, 160, NA),
is_sum = c(FALSE, FALSE, FALSE, FALSE, FALSE, TRUE)
)
ma$or_ci <- sprintf("%.2f (%.2f, %.2f)", ma$est, ma$lo, ma$hi)
forrest(
ma,
estimate = "est",
lower = "lo",
upper = "hi",
label = "study",
is_summary = "is_sum",
weight = "weight",
log_scale = TRUE,
ref_line = 1,
header = "Study",
cols = c("OR (95% CI)" = "or_ci"),
widths = c(2.2, 4, 2.5),
xlab = "Odds ratio (95% CI)"
)
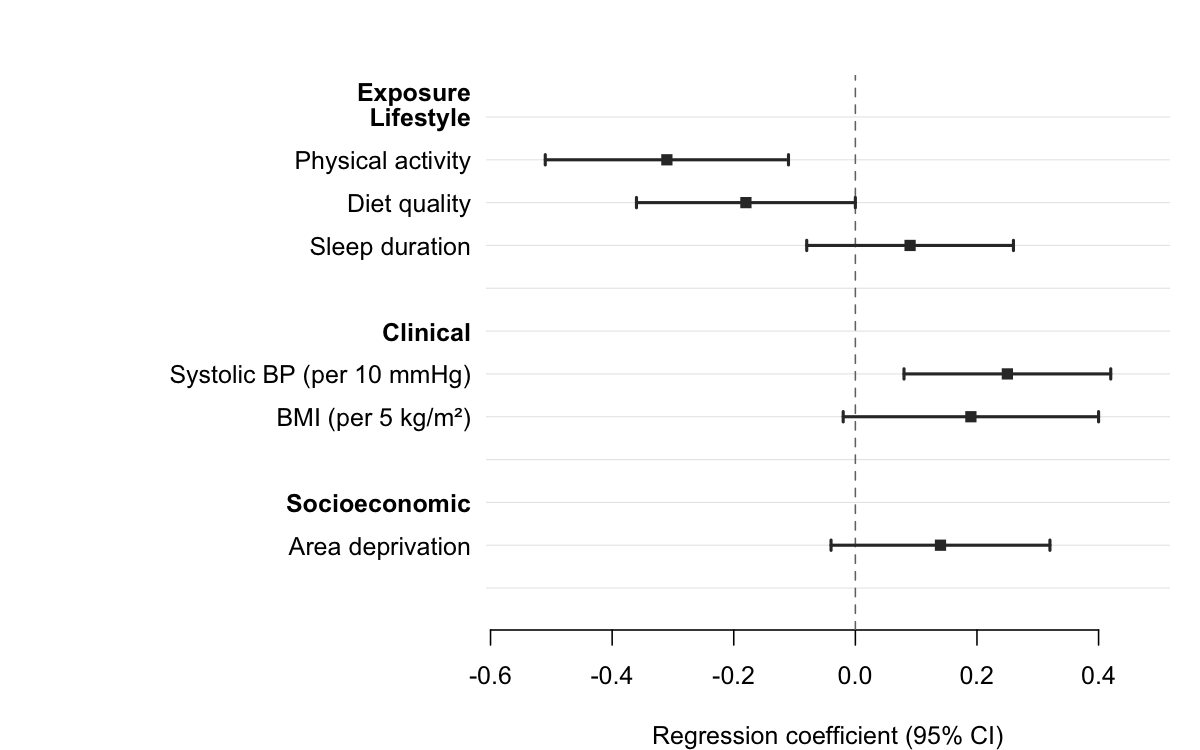
Pass a column name to section to group rows under
automatic bold headers. forrest() inserts a header wherever
the section value changes, indents the row labels, and adds a blank
spacer after each group — no manual data manipulation required.
subgroups <- data.frame(
domain = c("Lifestyle", "Lifestyle", "Lifestyle",
"Clinical", "Clinical",
"Socioeconomic"),
exposure = c("Physical activity", "Diet quality", "Sleep duration",
"Systolic BP (per 10 mmHg)", "BMI (per 5 kg/m\u00b2)",
"Area deprivation"),
est = c(-0.31, -0.18, 0.09, 0.25, 0.19, 0.14),
lo = c(-0.51, -0.36, -0.08, 0.08, -0.02, -0.04),
hi = c(-0.11, -0.00, 0.26, 0.42, 0.40, 0.32)
)
forrest(
subgroups,
estimate = "est",
lower = "lo",
upper = "hi",
label = "exposure",
section = "domain",
header = "Exposure",
ref_line = 0,
xlab = "Regression coefficient (95% CI)"
)
Use broom::tidy() to
extract parameters from a fitted model and pass the result directly to
forrest(). The cols argument adds a formatted
text column alongside the plot; header labels the row-label
panel. See the Regression
models vignette for progressive adjustment, multiple outcomes,
dose-response, and causal inference (g-computation, IPW) examples.
set.seed(1)
n <- 300
dat <- data.frame(
sbp = 120 + rnorm(n, sd = 15),
age = runif(n, 30, 70),
female = rbinom(n, 1, 0.5),
bmi = rnorm(n, 26, 4),
smoker = rbinom(n, 1, 0.2)
)
dat$sbp <- dat$sbp + 0.3 * dat$age - 4 * dat$female +
0.5 * dat$bmi - 2 * dat$smoker + rnorm(n, sd = 6)
fit <- lm(sbp ~ age + female + bmi + smoker, data = dat)
coefs <- broom::tidy(fit, conf.int = TRUE)
coefs <- coefs[coefs$term != "(Intercept)", ]
coefs$term <- c("Age (per 1 y)", "Female sex",
"BMI (per 1 kg/m²)", "Current smoker")
coefs$coef_ci <- sprintf(
"%.2f (%.2f, %.2f)",
coefs$estimate, coefs$conf.low, coefs$conf.high
)
forrest(
coefs,
estimate = "estimate",
lower = "conf.low",
upper = "conf.high",
label = "term",
header = "Predictor",
cols = c("Coef (95% CI)" = "coef_ci"),
widths = c(3, 4, 2.5),
xlab = "Regression coefficient (95% CI)",
stripe = TRUE
)
| Feature | forrest | forestplot | forestploter | ggforestplot |
|---|---|---|---|---|
| Minimal dependencies | ✅ | ❌ | ❌ | ❌ |
| General (not meta-analysis only) | ✅ | ✅ | ✅ | ⚠️ |
| Study weight sizing | ✅ | ✅ | ✅ | ❌ |
| Auto section headers from grouping column | ✅ | ⚠️ | ❌ | ❌ |
| Two-level nested section hierarchy | ✅ | ❌ | ❌ | ❌ |
| Summary diamonds | ✅ | ✅ | ✅ | ❌ |
| Text columns | ✅ | ✅ | ✅ | ❌ |
| Alternating row stripes | ✅ | ✅ | ✅ | ✅ |
| CI clipping with arrows | ✅ | ✅ | ⚠️ | ❌ |
| Group colouring + legend | ✅ | ✅ | ✅ | ✅ |
| Log-scale axis | ✅ | ✅ | ✅ | ✅ |
| Export helper | ✅ | ❌ | ⚠️ | ❌ |
| data.table support | ✅ | ❌ | ❌ | ❌ |
| Actively maintained | ✅ | ✅ | ✅ | ❌ |